|
Size: 13043
Comment: conda-init activation text
|
Size: 13207
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 8: | Line 8: |
| '''''Note: If you need to install EMAN2 within an existing Anaconda installation, please follow the source installation instructions instead, beginning at step 5. |
Binary Installation Instructions v2.91
Contents
Note: If you need to install EMAN2 within an existing Anaconda installation, please follow the source installation instructions instead, beginning at step 5.
Download older EMAN2 versions, instructions for older versions are here. Official releases are in reverse chronological order. The Continuous Build is rebuilt daily and any time a developer makes a change they think users should have access to. It is normally reasonably stable, and will contain the latest pre-publication features. Alternatively, the highest numbered version will contain a stable and tested release.
The neural network code in EMAN2 works best on GPUs, which are available only on the Linux installations. It can still run on Mac, but will be quite slow. EMAN2 initialization will add a block of code to your Mac users: If you don't understand what the .profile instructions are talking about, this may help: https://stackoverflow.com/questions/7501678/set-environment-variables-on-mac-os-x-lion Test: Run these programs to see if the install worked:
Linux users: The new neural network based routines are much faster running on a GPU. If you have an NVidia graphics card, see ''Using the GPU'' section below. Mac users (Big Sur): Big Sur (MacOS 11) includes new security features which require you to give software permission to access specific folders. This can cause problems for many open-source software packages launched from the command line. The easiest solution is Preferences -> Security & Privacy -> Privacy -> Full Disk Access, and give Terminal full access. This will not solve every problem, and you need to be aware of potential security risks in doing this, but for most scientific users it is a good solution. Mac users (Apple Silicon, M1): Anaconda does not yet fully support native M1 software, so you will need to install the normal Mac build. The installer will say that you appear not to be running on a 64 bit machine, and ask if you wish to install anyway. Say YES. Performance is excellent even though it isn't a native build. Check continuous build sections, Notes and Troubleshooting, which may contain more up-to-date info that may be relevant here.
If you get an error like the one below, dependency installation step might have failed. Run the dependency installation manually. Specifically, if only the last command fails and you are using an Nvidia graphics card, it is likely caused by a graphics card driver incompatibility. Updating the Nvidia driver usually fixes the problem. On recent Ubuntu systems, running If this causes issues for other software, you can type conda deactivate and EMAN2 commands will no longer work (but any software that doesn't like Anaconda will work). Alternatively, you can type conda config --set auto_activate_base False, which will prevent Anaconda from being activated automatically when you open new shells. In that case you will need to do a conda activate before running EMAN2/SPARX/SPHIRE commands. Check continuous build sections, Notes and Troubleshooting, which may contain more up-to-date info that may be relevant here.
While we do provide 64 bit Windows binaries for EMAN2, some functionality may be broken, so we strongly recommend using the #Windows-WSL option below if at all possible. Notes for native Windows version:
Download Download EMAN2.91. Select Installation Type: Just Me Choose Installation Location: Select a location with NO space in path Advanced Installation Options: Don't add EMAN2 to PATH environment variable. Open Anaconda Prompt by clicking Windows Start Menu -> Anaconda3 (64-bit) -> Anaconda Prompt or Start Menu -> Anaconda3 (64-bit) -> Anaconda Prompt(installation-folder-name). In most cases you will want to install: Python Launcher (launchwin-1.0.1.6.msi).
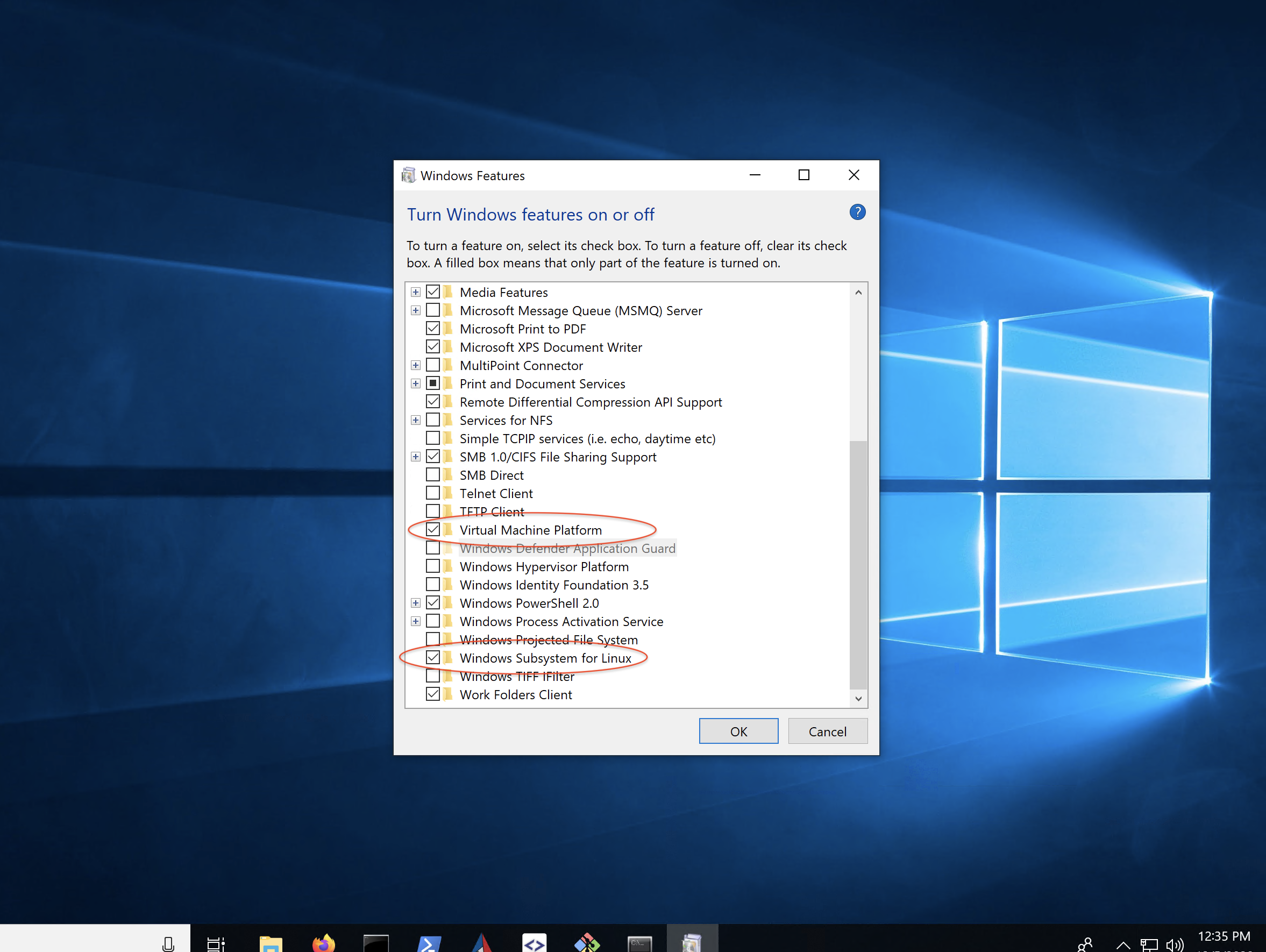
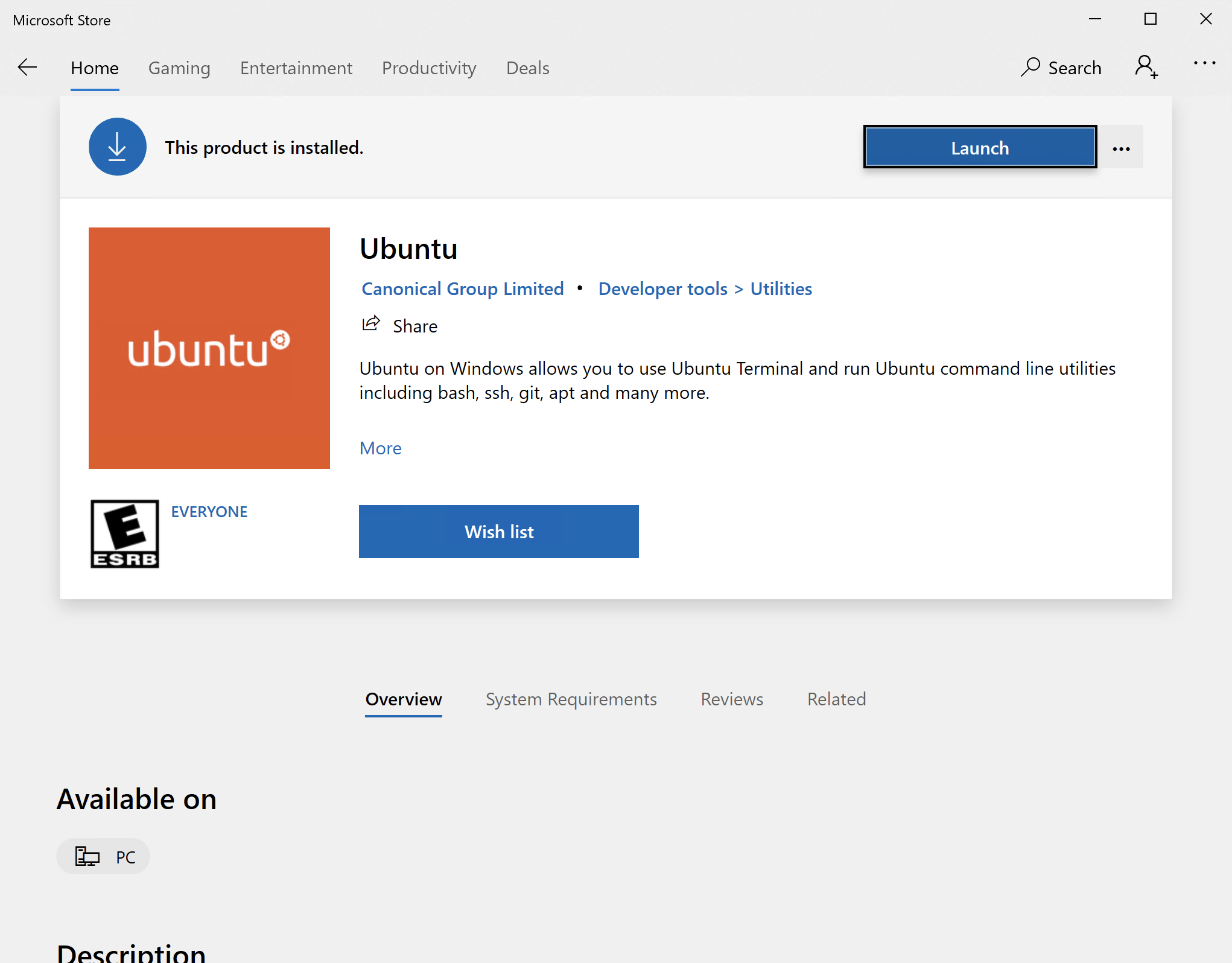
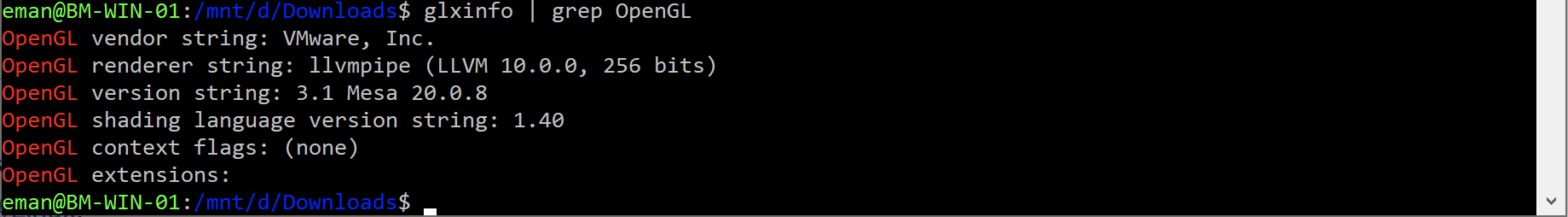
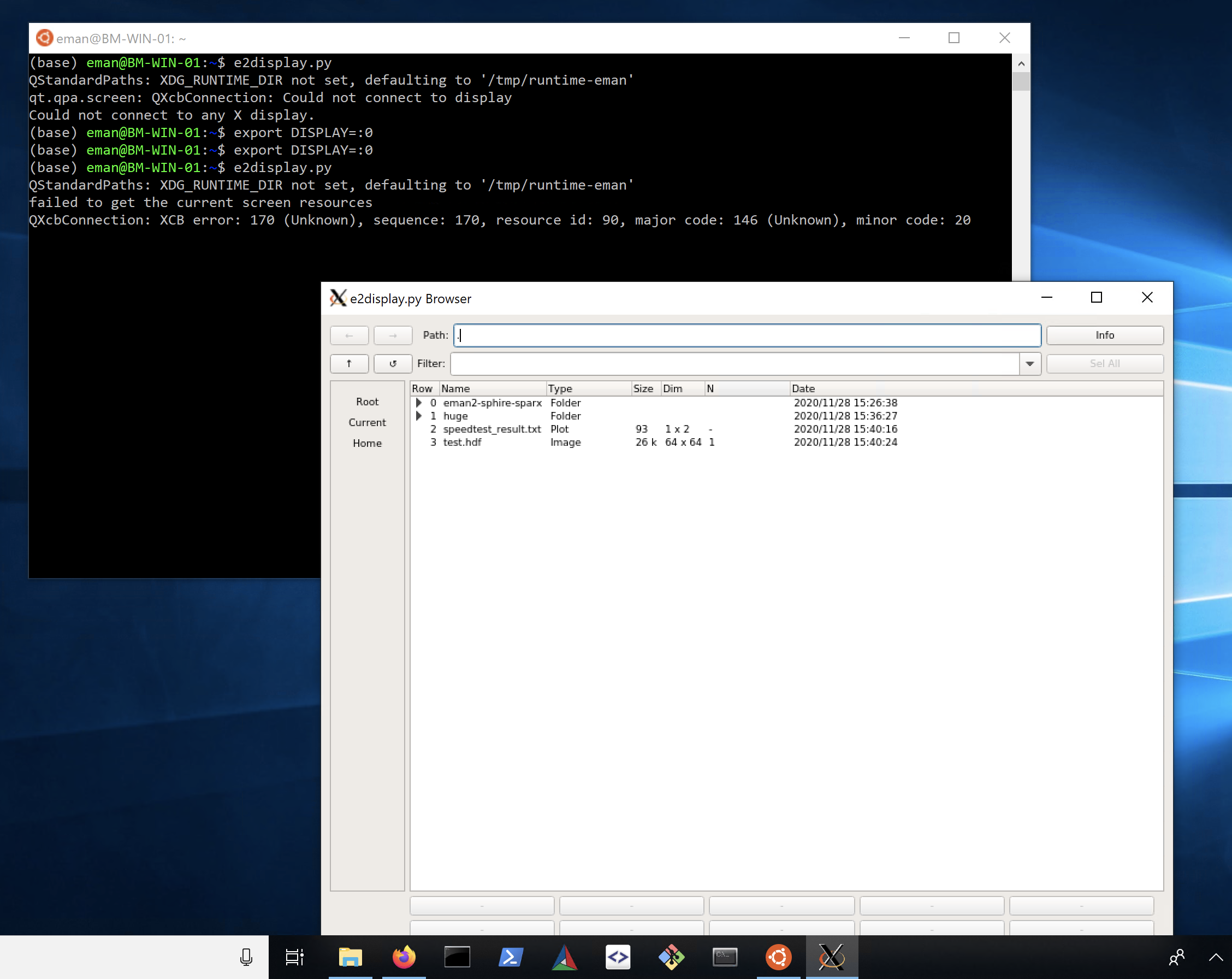
Windows 10 includes an embedded Ubuntu Linux environment. It is possible to run the EMAN2 Linux binaries within this Win10 environment, but you will need to install some additional dependencies to do so. Also, you will effectively be running at a Linux command prompt, so you will have to become a bit familiar with Linux to do this, but it does avoid installing an additional operating system on your machine. Install Xming X Server for Windows. Download and install LINUX binary, not windows binary! Download EMAN2.91, #Linux. Make sure to follow the instructions for shell initialization using conda-init.
Currently, the GPU is only used for neural network operations in tomogram annotation and in particle picking. It provides a ~10 fold or more speed up in neural network training. The new GPU developments are currently based on TensorFlow. From about 2006-2012 EMAN2 had its own internal CUDA code, which could be compiled into the C++ library. This has been deprecated, and likely no longer works, though the code is still present. We are working on a new GPU support strategy moving forward. Many machines will have CUDA installed already, and if CUDA is an appropriate version, this should work fine with the TensorFlow version shipped with EMAN2. However, if you are running newer versions of CUDA there may be problems. You can test compatibility quickly with: If this command does not return an error, then you should be able to run deep learning software within EMAN2. If it does raise an error, then you will need to debug the problem: Download
Install
MacOS and Linux
Cleanup: If you have previously installed EMAN2: 1 bash <path-to-EMAN2-installer>
.profile/.bashrc file. After initialization open your .profile/.bashrc file and confirm that a block like the following has been added Notes
Troubleshooting and Tips
apt-get install nvidia-current works. On other systems, you may need to follow the installation guide from Nvidia. Traceback (most recent call last):
File "/.../.../eman2/bin/e2proc3d.py", line 35, in <module>
from past.utils import old_div
ModuleNotFoundError: No module named 'past'conda install eman-deps=25 -c cryoem -c defaults -c conda-forge
Windows
Native Win7/10 64 bit
e2version.py
e2speedtest.py
e2display.py
e2proc2d.py :64:64:1 test.hdf --process mask.sharp:outer_radius=24
Windows 10 - Linux/Bash shell (As of 12/15/2020)
Using the GPU (Linux only)
# Make sure you have your environment set to run EMAN2 programs
e2version.py
# The above command should work and return your current version. If it does, then run:
python -c "import tensorflow"
apt-get install nvidia-cuda-toolkit
conda remove tensorflow-gpu tensorflow-gpu-base
pip install cython
pip install tensorflow
# read any messages carefully, if there are errors you may need other installations
[dnn]
enabled=False
[gcc]
cxxflags = -Wno-narrowing
