Extra functions for EMAN2 tomography
- This page describes extra functionalities of EMAN2 tomography workflow. This tutorial is frequently updated, so it is better to have EMAN2 version as new as possible.
Map particles to tomograms
There is a simple tool to map the averaged structure to the determined position and orientation of each particle in a tomogram. Available after EMAN2.3. In versions after 05/23/2019, the function is moved to the Analysis and Visualization section in the GUI.
Subtomogram Averaging -> Map particles to tomograms
Set path to be one of the spt_XX folder (not the subtlt ones).
Set iter to be the iteration you want to use from the refinement.
- Browse for one tomogram you want to map the particles to.
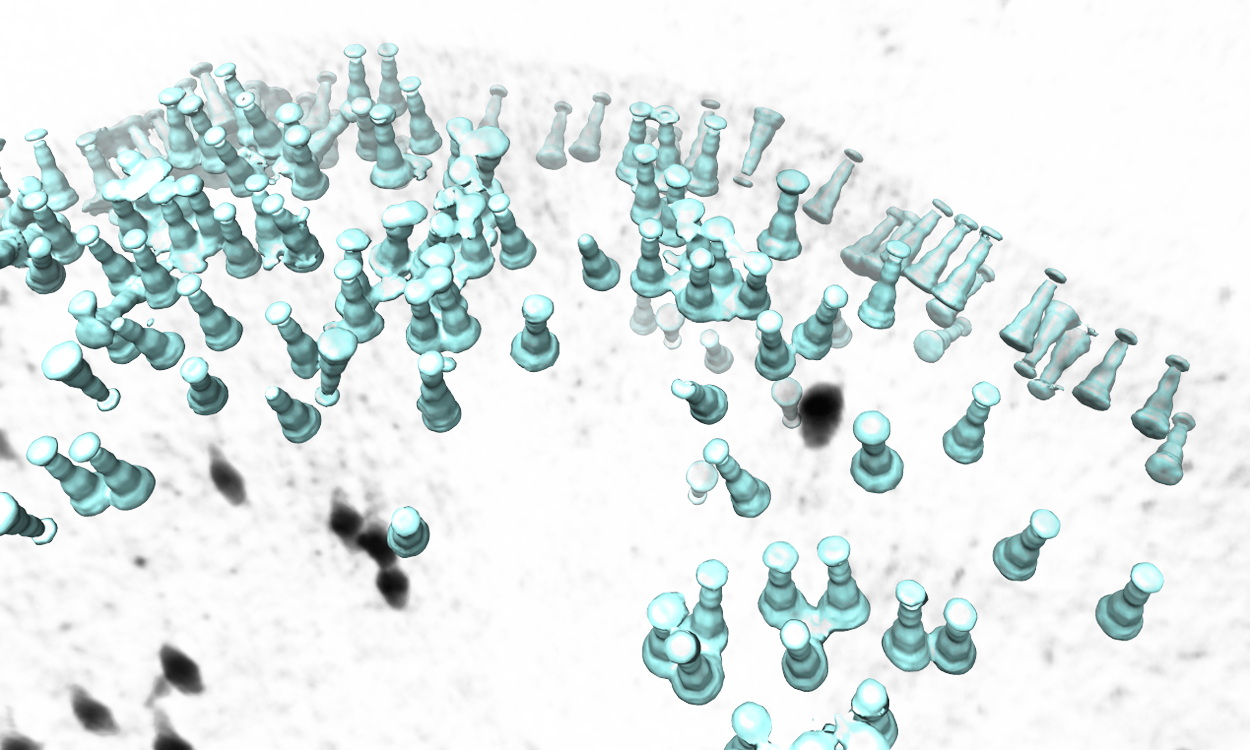
The program will then find all particles in the selected tomogram that are used in the refinement, map the averaged structure back, and produce a file called ptcls_in_tomo_xx_yy.hdf, where xx is the name of tomogram and yy is the number of iteration used. This is sometimes useful for objects in cellular environment but is quite uninteresting in this ribosome dataset. Image rendered with Chimera.