EMDash
EMDash is an EMEN2 client program for uploading images during a microscopy session. It includes tools for session management, commenting, viewing records, uploading, and watching data directories for new files.
Contents
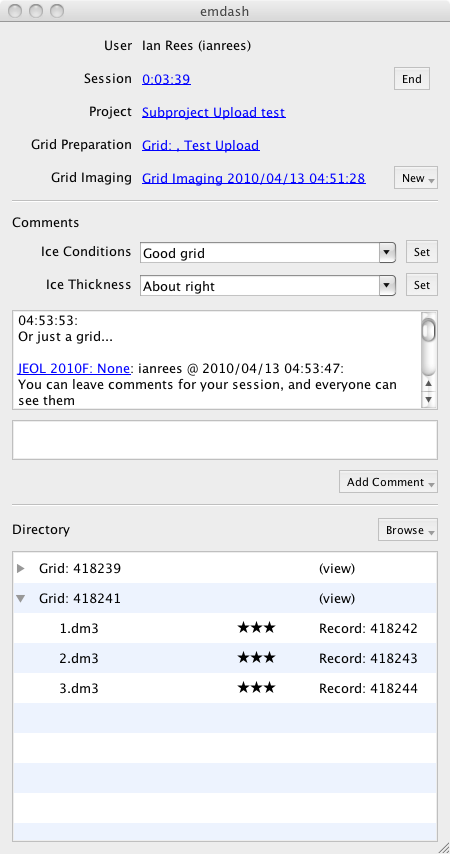
This is a screenshot of an active EMDash session. Each component in the interface will be described in the following sections.
Startup
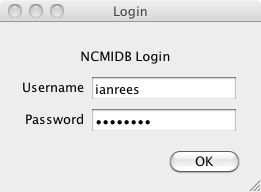
Start EMDash either in the System Tray in Windows, or like any other Python program. You will first be presented with a login screen:
New Grid Wizard
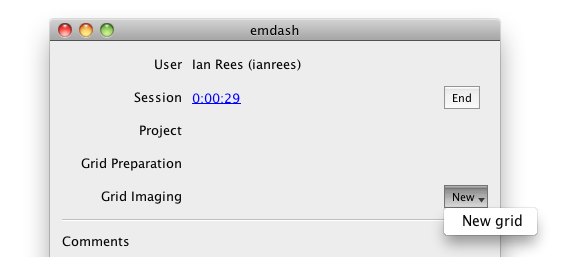
First, select the "New" dropdown to the right of "Grid Imaging," and select the option to create a new grid imaging session. You will be guided through a wizard that will ask you to select a Project or Subproject, enter the freezing details of the grid and any other preparation techniques used, and enter the basic overall parameters for this grid imaging session.
Page 1. Select a project
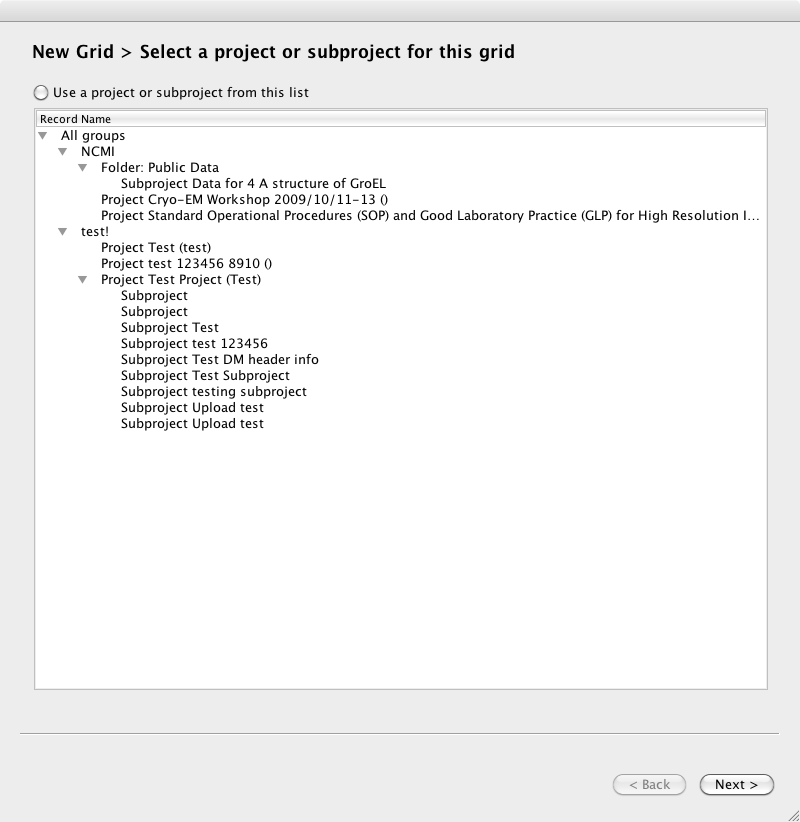
Page 2. Select a grid, or prompt to create a new grid. If you select an existing grid, you will skip the next screen.
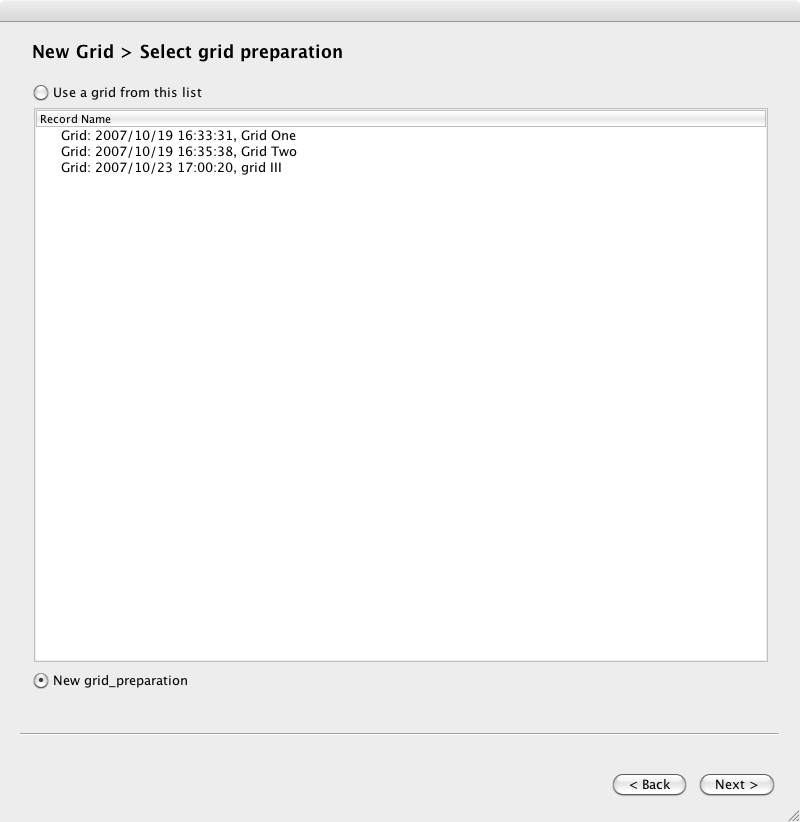
Page 3. Enter the grid preparation details
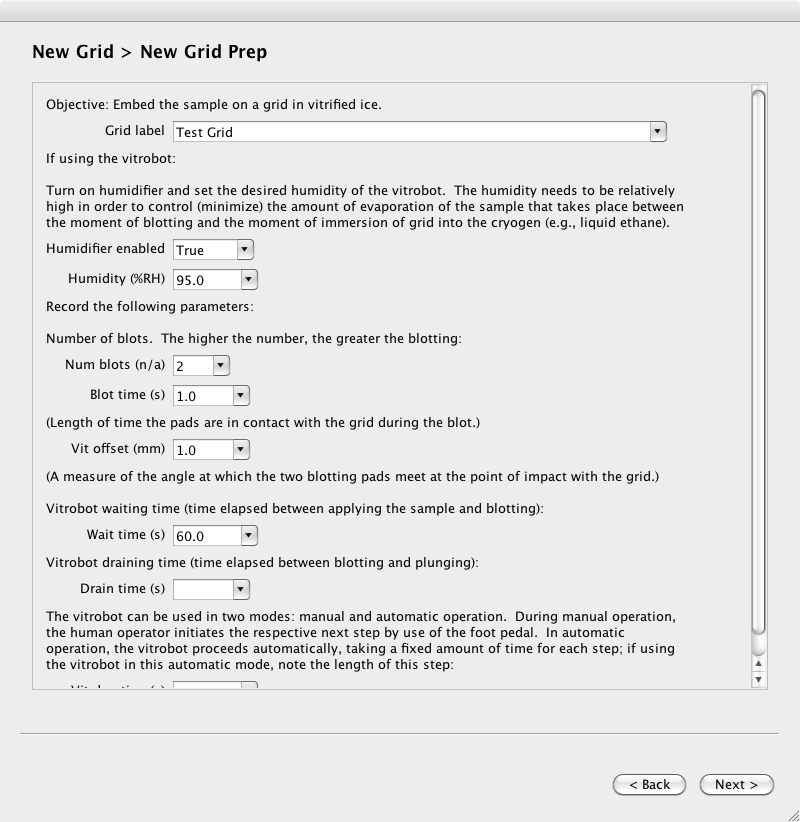
Page 4. Describe the purpose of the grid session, and provide basic microscope setup parameters.
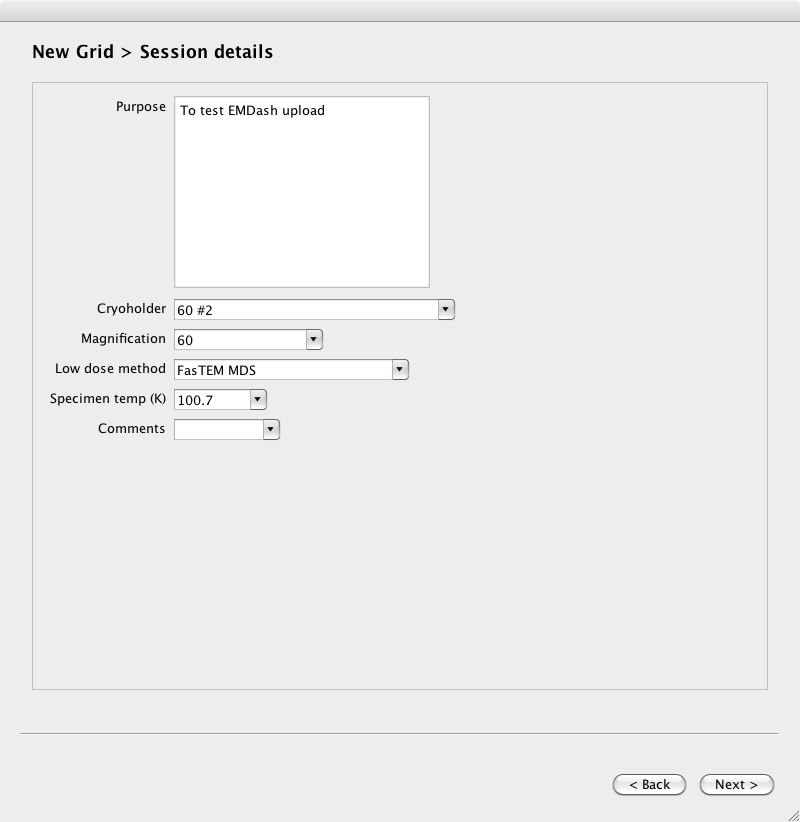
Selecting Files
EMDash will watch a directory for new files. New files will be added to the watch list, and CCD frames will be start to upload within seconds. To set the current watch directory, click the "Browse" button and select "Select directory." Use the normal file dialog window to find your data directory and confirm the dialog.
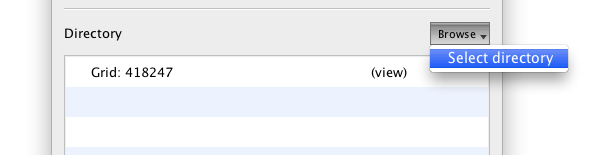
Any existing files will be picked up immediately, and others will be added as they are created.
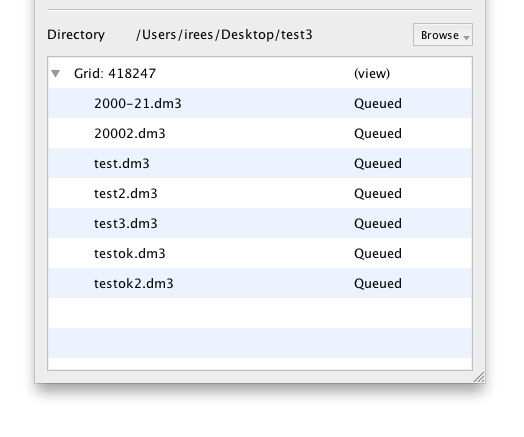
CCD Frames
Normal CCD frames will upload in the background as soon as they are closed in Digital Micrograph
Stack Files
Because stack files are written in append mode until SerielEM is finished with the entire tomography stack, you must manually tell EMDash when to begin the stack file upload. Click in the area labeled "Waiting" and select "Add to Queue." The stack will begin uploading immediately. There is a minor issue with status messages and progress being reported back to the GUI at the moment -- you will want to keep an eye on the console log to follow the progress of your stack file upload.
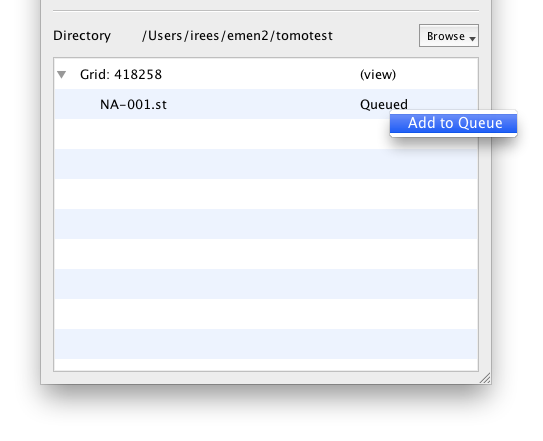
Be aware that, unlike CCD frames, you must have significant free space on the main hard disk available to upload stacks. EMDash, like emen2client, extracts all the slices and uploads each file individually. This was done for historical reasons, but since the file transfer capabilities of EMEN2 have improved, this may change in the future -- e.g. all stacks will be uploaded as-is and processed on the server.
Error Handling
EMDash will report any errors that occur during upload and automatically re-schedule any aborted file for upload. The most common events will be loss of network connectivity or temporary database unavailability (e.g. restarting.)
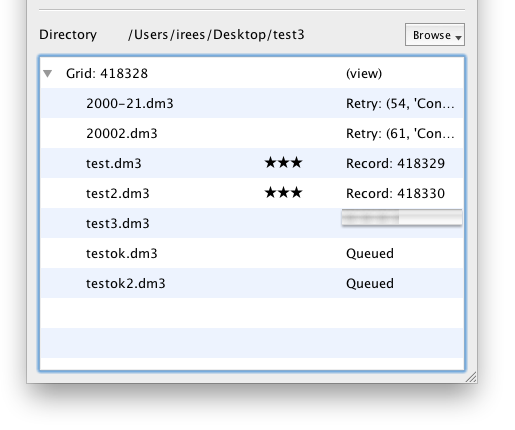
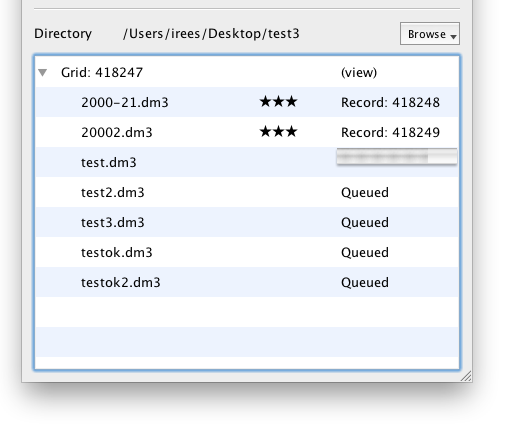
Rating
You can rate each image from Zero (Trash) to Five stars. After a file has been uploaded, the default rating of 3 stars will appear. You can click this to show an editor control. Click the star rating you wish to apply and it will be saved immediately to the database.
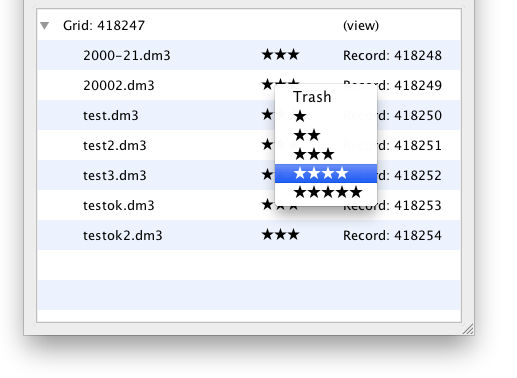
Note the actual rating system stored from 0-10 inside EMEN2 and EMAN2 for compatibility with existing convention; the five star range was chosen for simplicity. The stars are scaled by the following stars:rating; 0:0, 1:1, 2:3, 3:5, 4:7, 5:9.
Evaluating Your Session
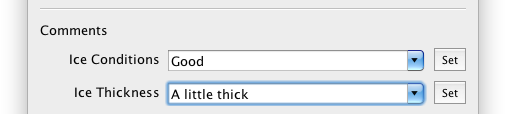
To describe the ice conditions and thickness in a particular grid, two special fields are provided in the comments area. Enter or select a value, and click "set" to save the value.
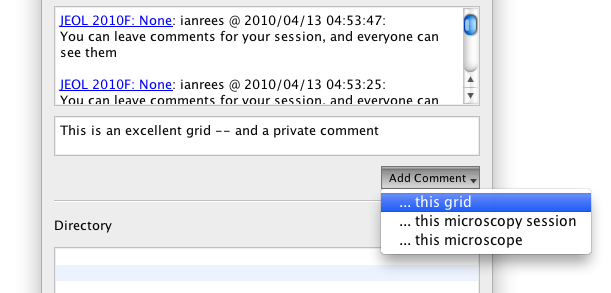
Adding Comments
You can attach comments to either your Microscopy session -- which is considered public -- or your Grid Imaging session -- which is restricted to members of the selected project.
Ending Your Session
The "End" button to the right of the "Session" timer will end your session and mark it as completed in the database. Any files that are currently queued for upload will continue to upload in the background as long as the machine is online. A warning dialog will be displayed to other microscope users to indicate this status. It will automatically close out and exit once the upload queue is finished.